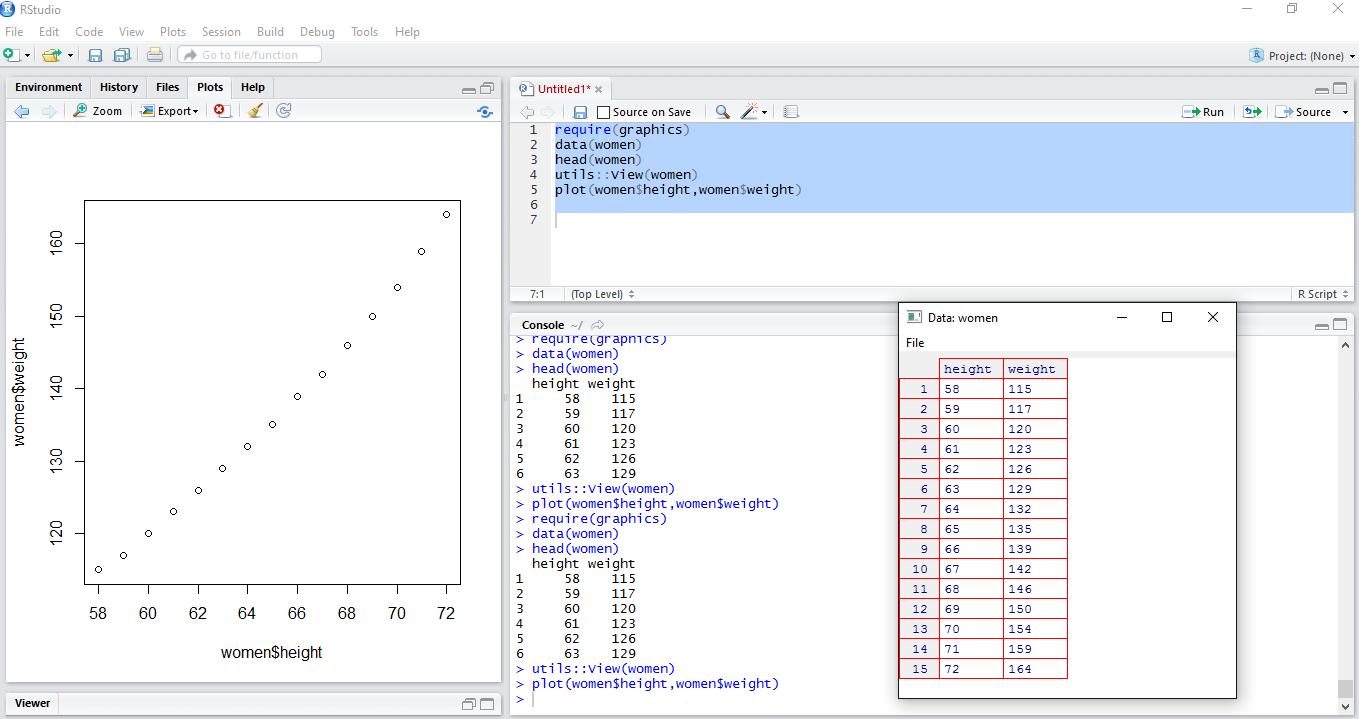
Our second R script is a simple two variable regression model which predicts womens height based upon weight.
Begin by creating another R script by selecting File | New File | R Script from the top navigation bar. If you create new scripts via File | New File | R Script often enough you might get Click Fatigue (uses three clicks), so you can also save a click by selecting the icon in the top left with the + sign:
Whichever way you choose , a new blank script window will appear with the name Untitled2.
Now paste the following code into the new script window:
require(graphics)
data(women)
lm_output <- lm(women$height ~ women$weight)
summary(lm_output)
prediction <- predict(lm_output)
error <- women$height-prediction
plot(women$height,error)
Press the Source icon to run the entire code. The display will change to something similar to what is displayed as follows:
Here are some notes and explanations for the script code that you have just run:
lm() function: This function runs a simple linear regression using the lm() function. This function predicts women's height based upon the value of their weight. In statistical parlance, you will be regressing height on weight. The line of code which accomplishes this is:
lm_output <- lm(women$height ~ women$weight)
- There are two operations that you will become very familiar with when running predictive models in R:
- The
~ operator: Also called the tilde, this is a shorthand way for separating what you want to predict, with what you are using to predict. This is an expression in formula syntax. What you are predicting (the dependent or target variable) is usually on the left side of the formula, and the predictors (independent variables, features) are on the right side. In order to improve readability, the independent variable (weight) and dependent variable (height) are specified using $ notation which specifies the object name, $, and then the dataframe column. So womens height is referenced as women$height and womens weight is referenced as women$weight. Alternatively, you can use the attach command, and then refer to these columns only by specifying the names height and weight. For example, the following code would achieve the same results:
attach(women)
lm_output <- lm(height ~ weight)- The
<- operator: Also called the assignment operator. This common statement assigns whatever expressions are evaluated on the right side of the assignment operator to the object specified on the left side of the operator. This will always create or replace a new object that you can further display or manipulate. In this case, we will be creating a new object called lm_output, which is created using the function lm(), which creates a linear model based on the formula contained within the parentheses.
Note
Note that the execution of this line does not produce any displayed output. You can see whether the line was executed by checking the console. If there is any problem with running the line (or any line for that matter), you will see an error message in the console.
summary(lm_output): The following statement displays some important summary information about the object lm_output and writes to output to the R Console as pictured previously:
summary(lm_output)
- The results will appear in the
Console window as pictured in the previous figure. Just to keep thing a little bit simpler for now, I will just show the first few lines of the output, and underline what you should be looking at. Do not be discouraged by the amount of output produced.
- Look at the lines marked
Intercept and women$weight which appear under the coefficients line in the console.
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 25.723456 1.043746 24.64 2.68e-12 ***
women$weight 0.287249 0.007588 37.85 1.09e-14 ***- The
Estimate column illustrates the linear regression formula needed to derive height from weight. We can actually use these numbers along with a calculator to determine the prediction ourselves. For our example the output tells us that we should perform the following steps for all of the observations in our dataframe in order to obtain the prediction for height. We will obviously not want to do all of the observations (R will do that via the following predict() function), but we will illustrate the calculation for 1 data point:
- Take the weight value for each observation. Lets take the weight of the first woman which is 115 lbs.
- Then,multiply weight by 0.2872 . That is the number that is listed under Estimate for womens$weight. Multiplying 115 lbs. by 0.2872 yield 33.028
- Then add 25.7235 which is the estimate of the (intercept) row. That will yield a prediction of 58.75 inches.
- If you do not have a calculator handy, the calculation is easily done in calculator mode via the R Console, by typing the following:
To predict the value for all of the values we will use a function called predict(). This function reads each input (independent) variable and then predicts a target (dependent) variable based on the linear regression equation. In the code we have assigned the output of this function to a new object named prediction.
Switch over to the console area, and type prediction, then Enter, to see the predicted values for the 15 women. The following should appear in the console.
> prediction
1 2 3 4 5 6 7
58.75712 59.33162 60.19336 61.05511 61.91686 62.77861 63.64035
8 9 10 11 12 13 14
64.50210 65.65110 66.51285 67.66184 68.81084 69.95984 71.39608
15
72.83233 Notice that the value of the first prediction is very close to what you just calculated by hand. The difference is due to rounding error.
Examining the prediction errors
Another R object produced by our linear regression is the error object. The error object is a vector that was computed by taking the difference between the predicted value of height and the actual height. These values are also known as the residual errors, or just residuals.
error <- women$height-prediction
Since the error object is a vector, you cannot use the nrow() function to get its size. But you can use the length() function:
>length(error)
[1] 15
In all of the previous cases, the counts all total 15, so all is good. If we want to see the raw data, predictions, and the prediction errors for all of the data, we can use the cbind() function (Column bind) to concatenate all three of those values, and display as a simple table.
At the console enter the follow cbind command:
> cbind(height=women$height,PredictedHeight=prediction,ErrorInPrediction=error)
height PredictedHeight ErrorInPrediction
1 58 58.75712 -0.75711680
2 59 59.33162 -0.33161526
3 60 60.19336 -0.19336294
4 61 61.05511 -0.05511062
5 62 61.91686 0.08314170
6 63 62.77861 0.22139402
7 64 63.64035 0.35964634
8 65 64.50210 0.49789866
9 66 65.65110 0.34890175
10 67 66.51285 0.48715407
11 68 67.66184 0.33815716
12 69 68.81084 0.18916026
13 70 69.95984 0.04016335
14 71 71.39608 -0.39608278
15 72 72.83233 -0.83232892
From the preceding output, we can see that there are a total 15 predictions. If you compare the ErrorInPrediction with the error plot shown previously, you can see that for this very simple model, the prediction errors are much larger for extreme values in height (shaded values).
Just to verify that we have one for each of our original observations we will use the nrow() function to count the number of rows.
At the command prompt in the console area, enter the command:
nrow(women)
The following should appear:
>nrow(women)
[1] 15
Refer back to the seventh line of code in the original script: plot(women$height,error) plots the predicted height versus the errors. It shows how much the prediction was off from the original value. You can see that the errors show a non-random pattern.
After you are done, save the file using File | File Save, navigate to the PracticalPredictiveAnalytics/R folder that was created, and name it Chapter1_LinearRegression.
 Argentina
Argentina
 Australia
Australia
 Austria
Austria
 Belgium
Belgium
 Brazil
Brazil
 Bulgaria
Bulgaria
 Canada
Canada
 Chile
Chile
 Colombia
Colombia
 Cyprus
Cyprus
 Czechia
Czechia
 Denmark
Denmark
 Ecuador
Ecuador
 Egypt
Egypt
 Estonia
Estonia
 Finland
Finland
 France
France
 Germany
Germany
 Great Britain
Great Britain
 Greece
Greece
 Hungary
Hungary
 India
India
 Indonesia
Indonesia
 Ireland
Ireland
 Italy
Italy
 Japan
Japan
 Latvia
Latvia
 Lithuania
Lithuania
 Luxembourg
Luxembourg
 Malaysia
Malaysia
 Malta
Malta
 Mexico
Mexico
 Netherlands
Netherlands
 New Zealand
New Zealand
 Norway
Norway
 Philippines
Philippines
 Poland
Poland
 Portugal
Portugal
 Romania
Romania
 Russia
Russia
 Singapore
Singapore
 Slovakia
Slovakia
 Slovenia
Slovenia
 South Africa
South Africa
 South Korea
South Korea
 Spain
Spain
 Sweden
Sweden
 Switzerland
Switzerland
 Taiwan
Taiwan
 Thailand
Thailand
 Turkey
Turkey
 Ukraine
Ukraine
 United States
United States