In this chapter, you will learn how to apply unsupervised learning techniques to identify patterns and structure within datasets.
Unsupervised learning techniques are a valuable set of tools for exploratory analysis. They bring out patterns and structure within datasets, which yield information that may be informative in itself or serve as a guide to further analysis. It's critical to have a solid set of unsupervised learning tools that you can apply to help break up unfamiliar or complex datasets into actionable information.
We'll begin by reviewing Principal Component Analysis (PCA), a fundamental data manipulation technique with a range of dimensionality reduction applications. Next, we will discuss k-means clustering, a widely-used and approachable unsupervised learning technique. Then, we will discuss Kohenen's Self-Organizing Map (SOM), a method of topological clustering that enables the projection of complex datasets into two dimensions.
Throughout the chapter, we will spend some time discussing how to effectively apply these techniques to make high-dimensional datasets readily accessible. We will use the UCI Handwritten Digits dataset to demonstrate technical applications of each algorithm. In the course of discussing and applying each technique, we will review practical applications and methodological questions, particularly regarding how to calibrate and validate each technique as well as which performance measures are valid. To recap, then, we will be covering the following topics in order:
Principal component analysis
k-means clustering
Self-organizing maps
In order to work effectively with high-dimensional datasets, it is important to have a set of techniques that can reduce this dimensionality down to manageable levels. The advantages of this dimensionality reduction include the ability to plot multivariate data in two dimensions, capture the majority of a dataset's informational content within a minimal number of features, and, in some contexts, identify collinear model components.
Note
For those in need of a refresher, collinearity in a machine learning context refers to model features that share an approximately linear relationship. For reasons that will likely be obvious, these features tend to be unhelpful as the related features are unlikely to add information mutually that either one provides independently. Moreover, collinear features may emphasize local minima or other false leads.
Probably the most widely-used dimensionality reduction technique today is PCA. As we'll be applying PCA in multiple contexts throughout this book, it's appropriate for us to review the technique, understand the theory behind it, and write Python code to effectively apply it.
PCA is a powerful decomposition technique; it allows one to break down a highly multivariate dataset into a set of orthogonal components. When taken together in sufficient number, these components can explain almost all of the dataset's variance. In essence, these components deliver an abbreviated description of the dataset. PCA has a broad set of applications and its extensive utility makes it well worth our time to cover.
Note
Note the slightly cautious phrasing here—a given set of components of length less than the number of variables in the original dataset will almost always lose some amount of the information content within the source dataset. This lossiness is typically minimal, given enough components, but in cases where small numbers of principal components are composed from very high-dimensional datasets, there may be substantial lossiness. As such, when performing PCA, it is always appropriate to consider how many components will be necessary to effectively model the dataset in question.
PCA works by successively identifying the axis of greatest variance in a dataset (the principal components). It does this as follows:
Let's unpack these concepts briefly:
Covariance is effectively variance applied to multiple dimensions; it is the variance between two or more variables. While a single value can capture the variance in one dimension or variable, it is necessary to use a 2 x 2 matrix to capture the covariance between two variables, a 3 x 3 matrix to capture the covariance between three variables, and so on. So the first step in PCA is to calculate this covariance matrix.
An Eigenvector is a vector that is specific to a dataset and linear transformation. Specifically, it is the vector that does not change in direction before and after the transformation is performed. To get a better feeling for how this works, imagine that you're holding a rubber band, straight, between both hands. Let's say you stretch the band out until it is taut between your hands. The eigenvector is the vector that did not change direction between before the stretch and during it; in this case, it's the vector running directly through the center of the band from one hand to the other.
Orthogonalization is the process of finding two vectors that are orthogonal (at right angles) to one another. In an n-dimensional data space, the process of orthogonalization takes a set of vectors and yields a set of orthogonal vectors.
Orthonormalization is an orthogonalization process that also normalizes the product.
Eigenvalue (roughly corresponding to the length of the eigenvector) is used to calculate the proportion of variance represented by each eigenvector. This is done by dividing the eigenvalue for each eigenvector by the sum of eigenvalues for all eigenvectors.
In summary, the covariance matrix is used to calculate Eigenvectors. An orthonormalization process is undertaken that produces orthogonal, normalized vectors from the Eigenvectors. The eigenvector with the greatest eigenvalue is the first principal component with successive components having smaller eigenvalues. In this way, the PCA algorithm has the effect of taking a dataset and transforming it into a new, lower-dimensional coordinate system.
Now that we've reviewed the PCA algorithm at a high level, we're going to jump straight in and apply PCA to a key Python dataset—the UCI handwritten digits dataset, distributed as part of
scikit-learn.
This dataset is composed of 1,797 instances of handwritten digits gathered from 44 different writers. The input (pressure and location) from these authors' writing is resampled twice across an 8 x 8 grid so as to yield maps of the kind shown in the following image:
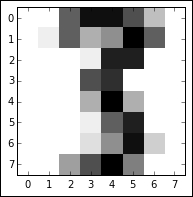
These maps can be transformed into feature vectors of length 64, which are then readily usable as analysis input. With an input dataset of 64 features, there is an immediate appeal to using a technique like PCA to reduce the set of variables to a manageable amount. As it currently stands, we cannot effectively explore the dataset with exploratory visualization!
We will begin applying PCA to the handwritten digits dataset with the following code:
import numpy as np from sklearn.datasets import load_digits import matplotlib.pyplot as plt from sklearn.decomposition import PCA from sklearn.preprocessing import scale from sklearn.lda import LDA import matplotlib.cm as cm digits = load_digits() data = digits.data n_samples, n_features = data.shape n_digits = len(np.unique(digits.target)) labels = digits.target
This code does several things for us:
First, it loads up a set of necessary libraries, including
numpy, a set of components from scikit-learn, including thedigitsdataset itself, PCA and data scaling functions, and the plotting capability of matplotlib.The code then begins preparing the
digitsdataset. It does several things in order:First, it loads the dataset before creating helpful variables
The
datavariable is created for subsequent use, and the number of distinctdigitsin thetargetvector (0 through to 9, son_digits = 10) is saved as a variable that we can easily access for subsequent analysisThe
targetvector is also saved as labels for later useAll of this variable creation is intended to simplify subsequent analysis
With the dataset ready, we can initialize our PCA algorithm and apply it to the dataset:
pca = PCA(n_components=10) data_r = pca.fit(data).transform(data) print('explained variance ratio (first two components): %s' % str(pca.explained_variance_ratio_)) print('sum of explained variance (first two components): %s' % str(sum(pca.explained_variance_ratio_)))This code outputs the variance explained by each of the first ten principal components ordered by explanatory power.
In the case of this set of 10 principal components, they collectively explain 0.589 of the overall dataset variance. This isn't actually too bad, considering that it's a reduction from 64 variables to 10 components. It does, however, illustrate the potential lossiness of PCA. The key question, though, is whether this reduced set of components makes subsequent analysis or classification easier to achieve; that is, whether many of the remaining components contained variance that disrupts classification attempts.
Having created a data_r object containing the output of pca performed over the digits dataset, let's visualize the output. To do so, we'll first create a vector of colors for class coloration. We then simply create a scatterplot with colorized classes:
X = np.arange(10)
ys = [i+x+(i*x)**2 for i in range(10)]
plt.figure()
colors = cm.rainbow(np.linspace(0, 1, len(ys)))
for c, i target_name in zip(colors, [1,2,3,4,5,6,7,8,9,10], labels):
plt.scatter(data_r[labels == I, 0], data_r[labels == I, 1],
c=c, alpha = 0.4)
plt.legend()
plt.title('Scatterplot of Points plotted in first \n'
'10 Principal Components')
plt.show()The resulting scatterplot looks as follows:
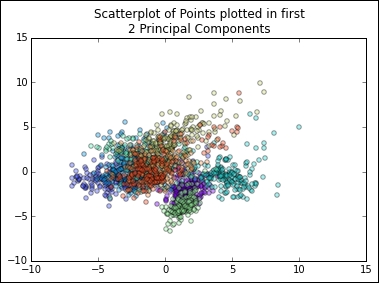
This plot shows us that, while there is some separation between classes in the first two principal components, it may be tricky to classify highly accurately with this dataset. However, classes do appear to be clustered and we may be able to get reasonably good results by employing a clustering analysis. In this way, PCA has given us some insight into how the dataset is structured and has informed our subsequent analysis.
At this point, let's take this insight and move on to examine clustering by the application of the k-means clustering algorithm.
In the previous section, you learned that unsupervised machine learning algorithms are used to extract key structural or information content from large, possibly complex datasets. These algorithms do so with little or no manual input and function without the need for training data (sets of labeled explanatory and response variables needed to train an algorithm in order to recognize the desired classification boundaries). This means that unsupervised algorithms are effective tools to generate information about the structure and content of new or unfamiliar datasets. They allow the analyst to build a strong understanding in a fraction of the time.
Clustering is probably the archetypal unsupervised learning technique for several reasons.
A lot of development time has been sunk into optimizing clustering algorithms, with efficient implementations available in most data science languages including Python.
Clustering algorithms tend to be very fast, with smoothed implementations running in polynomial time. This makes it uncomplicated to run multiple clustering configurations, even over large datasets. Scalable clustering implementations also exist that parallelize the algorithm to run over TB-scale datasets.
Clustering algorithms are frequently easily understood and their operation is thus easy to explain if necessary.
The most popular clustering algorithm is k-means; this algorithm forms k-many clusters by first randomly initiating the clusters as k-many points in the data space. Each of these points is the mean of a cluster. An iterative process then occurs, running as follows:
Each point is assigned to a cluster based on the least (within cluster) sum of squares, which is intuitively the nearest mean.
The center (centroid) of each cluster becomes the new mean. This causes each of the means to shift.
Over enough iterations, the centroids move into positions that minimize a performance metric (the performance metric most commonly used is the "within cluster least sum of squares" measure). Once this measure is minimized, observations are no longer reassigned during iteration; at this point the algorithm has converged on a solution.
Now that we've reviewed the clustering algorithm, let's run through the code and see what clustering can do for us:
from time import time
import numpy as np
import matplotlib.pyplot as plt
np.random.seed()
digits = load_digits()
data = scale(digits.data)
n_samples, n_features = data.shape
n_digits = len(np.unique(digits.target))
labels = digits.target
sample_size = 300
print("n_digits: %d, \t n_samples %d, \t n_features %d"
% (n_digits, n_samples, n_features))
print(79 * '_')
print('% 9s' % 'init'' time inertia homo compl v-meas ARI AMI silhouette')
def bench_k_means(estimator, name, data):
t0 = time()
estimator.fit(data)
print('% 9s %.2fs %i %.3f %.3f %.3f %.3f %.3f %.3f'
% (name, (time() - t0), estimator.inertia_,
metrics.homogeneity_score(labels, estimator.labels_),
metrics.completeness_score(labels, estimator.labels_),
metrics.v_measure_score(labels, estimator.labels_),
metrics.adjusted_rand_score(labels, estimator.labels_),
metrics.silhouette_score(data, estimator.labels_,
metric='euclidean',
sample_size=sample_size)))Note
One critical difference between this code and the PCA code we saw previously is that this code begins by applying a scale function to the digits dataset. This function scales values in the dataset between 0 and 1. It's critically important to scale data wherever needed, either on a log scale or bound scale, so as to prevent the magnitude of different feature values to have disproportionately powerful effects on the dataset. The key to determining whether the data needs scaling at all (and what kind of scaling is needed, within which range, and so on) is very much tied to the shape and nature of the data. If the distribution of the data shows outliers or variation within a large range, it may be appropriate to apply log-scaling. Whether this is done manually through visualization and exploratory analysis techniques or through the use of summary statistics, decisions around scaling are tied to the data under inspection and the analysis techniques to be used. A further discussion of scaling decisions and considerations may be found in Chapter 7, Feature Engineering Part II.
Helpfully, scikit-learn uses the k-means++ algorithm by default, which improves over the original k-means algorithm in terms of both running time and success rate in avoiding poor clusterings.
The algorithm achieves this by running an initialization procedure to find cluster centroids that approximate minimal variance within classes.
You may have spotted from the preceding code that we're using a set of performance estimators to track how well our k-means application is performing. It isn't practical to measure the performance of a clustering algorithm based on a single correctness percentage or using the same performance measures that are commonly used with other algorithms. The definition of success for clustering algorithms is that they provide an interpretation of how input data is grouped that trades off between several factors, including class separation, in-group similarity, and cross-group difference.
The homogeneity score is a simple, zero-to-one-bounded measure of the degree to which clusters contain only assignments of a given class. A score of one indicates that all clusters contain measurements from a single class. This measure is complimented by the completeness score, which is a similarly bounded measure of the extent to which all members of a given class are assigned to the same cluster. As such, a completeness score and homogeneity score of one indicates a perfect clustering solution.
The validity measure (v-measure) is a harmonic mean of the homogeneity and completeness scores, which is exactly analogous to the F-measure for binary classification. In essence, it provides a single, 0-1-scaled value to monitor both homogeneity and completeness.
The Adjusted Rand Index (ARI) is a similarity measure that tracks the consensus between sets of assignments. As applied to clustering, it measures the consensus between the true, pre-existing observation labels and the labels predicted as an output of the clustering algorithm. The Rand index measures labeling similarity on a 0-1 bound scale, with one equaling perfect prediction labels.
The main challenge with all of the preceding performance measures as well as other similar measures (for example, Akaike's mutual information criterion) is that they require an understanding of the ground truth, that is, they require some or all of the data under inspection to be labeled. If labels do not exist and cannot be generated, these measures won't work. In practice, this is a pretty substantial drawback as very few datasets come prelabeled and the creation of labels can be time-consuming.
One option to measure the performance of a k-means clustering solution without labeled data is the Silhouette Coefficient. This is a measure of how well-defined the clusters within a model are. The Silhouette Coefficient for a given dataset is the mean of the coefficient for each sample, where this coefficient is calculated as follows:

The definitions of each term are as follows:
a: The mean distance between a sample and all other points in the same cluster
b: The mean distance between a sample and all other points in the next nearest cluster
This score is bounded between -1 and 1, with -1 indicating incorrect clustering, 1 indicating very dense clustering, and scores around 0 indicating overlapping clusters. This tends to fit our expectations of how a good clustering solution is composed.
In the case of the digits dataset, we can employ all of the performance measures described here. As such, we'll complete the preceding example by initializing our bench_k_means function over the digits dataset:
bench_k_means(KMeans(init='k-means++', n_clusters=n_digits, n_init=10), name="k-means++", data=data) print(79 * '_')
This yields the following output (note that the random seed means your results will vary from mine!):

Lets take a look at these results in more detail.
The Silhouette score at 0.123 is fairly low, but not surprisingly so, given that the handwritten digits data is inherently noisy and does tend to overlap. However, some of the other scores are not that impressive. The V-measure at 0.619 is reasonable, but in this case is held back by a poor homogeneity measure, suggesting that the cluster centroids did not resolve perfectly. Moreover, the ARI at 0.465 is not great.
Note
Let's put this in context. The worst case classification attempt, random assignment, would give at best 10% classification accuracy. All of our performance measures would be accordingly very low. While we're definitely doing a lot better than that, we're still trailing far behind the best computational classification attempts. As we'll see in Chapter 4, Convolutional Neural Networks, convolutional nets achieve results with extremely low classification errors on handwritten digit datasets. We're unlikely to achieve this level of accuracy with traditional k-means clustering!
All in all, it's reasonable to think that we could do better.
To give this another try, we'll apply an additional stage of processing. To learn how to do this, we'll apply PCA—the technique we previously walked through—to reduce the dimensionality of our input dataset. The code to achieve this is very simple, as follows:
pca = PCA(n_components=n_digits).fit(data) bench_k_means(KMeans(init=pca.components_, n_clusters=10), name="PCA-based", data=data)
This code simply applies PCA to the digits dataset, yielding as many principal components as there are classes (in this case, digits). It can be sensible to review the output of PCA before proceeding as the presence of any small principal components may suggest a dataset that contains collinearity or otherwise merits further inspection.
This instance of clustering shows noticeable improvement:

The V-measure and ARI have increased by approximately 0.08 points, with the V-measure reading a fairly respectable 0.693. The Silhouette Coefficient did not change significantly. Given the complexity and interclass overlap within the digits dataset, these are good results, particularly stemming from such a simple code addition!
Inspection of the digits dataset with clusters superimposed shows that some meaningful clusters appear to have been formed. It is also apparent from the following plot that actually detecting the character from the input feature vectors may be a challenging task:

The previous examples described how to apply k-means, walked through relevant code, showed how to plot the results of a clustering analysis, and identified appropriate performance metrics. However, when applying k-means to real-world datasets, there are some extra precautions that need to be taken, which we will discuss.
Another critical practical point is how to select an appropriate value for k. Initializing k-means clustering with a specific k value may not be harmful, but in many cases it is not clear initially how many clusters you might find or what values of k may be helpful.
We can rerun the preceding code for multiple values of k in a batch and look at the performance metrics, but this won't tell us which instance of k is most effectively capturing structure within the data. The risk is that as k increases, the Silhouette Coefficient or unexplained variance may decrease dramatically, without meaningful clusters being formed. The extreme case of this would be if k = o, where o is the number of observations in the sample; every point would have its own cluster, the Silhouette Coefficient would be low, but the results wouldn't be meaningful. There are, however, many less extreme cases in which overfitting may occur due to an overly high k value.
To mitigate this risk, it's advisable to use supporting techniques to motivate a selection of k. One useful technique in this context is the elbow method. The elbow method is a very simple technique; for each instance of k, plot the percentage of explained variance against k. This typically leads to a plot that frequently looks like a bent arm.
For the PCA-reduced dataset, this code looks like the following snippet:
import numpy as np from sklearn.cluster import KMeans from sklearn.datasets import load_digits from scipy.spatial.distance import cdist import matplotlib.pyplot as plt from sklearn.decomposition import PCA from sklearn.preprocessing import scale digits = load_digits() data = scale(digits.data) n_samples, n_features = data.shape n_digits = len(np.unique(digits.target)) labels = digits.target K = range(1,20) explainedvariance= [] for k in K: reduced_data = PCA(n_components=2).fit_transform(data) kmeans = KMeans(init = 'k-means++', n_clusters = k, n_init = k) kmeans.fit(reduced_data) explainedvariance.append(sum(np.min(cdist(reduced_data, kmeans.cluster_centers_, 'euclidean'), axis = 1))/data.shape[0]) plt.plot(K, meandistortions, 'bx-') plt.show()
This application of the elbow method takes the PCA reduction from the previous code sample and applies a test of the explained variance (specifically, a test of the variance within clusters). The result is output as a measure of unexplained variance for each value of k in the range specified. In this case, as we're using the digits dataset (which we know to have ten classes), the range specified was 1 to 20:

The elbow method involves selecting the value of k that maximizes explained variance while minimizing K; that is, the value of k at the crook of the elbow. The technical sense underlying this is that a minimal gain in explained variance at greater values of k is offset by the increasing risk of overfitting.
Elbow plots may be more or less pronounced and the elbow may not always be clearly identifiable. This example shows a more gradual progression than may be observable in other cases with other datasets. It's worth noting that, while we know the number of classes within the dataset to be ten, the elbow method starts to show diminishing returns on k increases almost immediately and the elbow is located at around five classes. This has a lot to do with the substantial overlap between classes, which we saw in previous plots. While there are ten classes, it becomes increasingly difficult to clearly identify more than five or so.
With this in mind, it's worth noting that the elbow method is intended for use as a heuristic rather than as some kind of objective principle. The use of PCA as a preprocess to improve clustering performance also tends to smooth the graph, delivering a more gradual curve than otherwise.
In addition to making use of the elbow method, it can be valuable to look at the clusters themselves, as we did earlier in the chapter, using PCA to reduce the dimensionality of the data. By plotting the dataset and projecting cluster assignation onto the data, it is sometimes very obvious when a k-means implementation has fitted to a local minima or has overfit the data. The following plot demonstrates extreme overfitting of our previous k-means clustering algorithm to the digits dataset, artificially prompted by using K = 150. In this example, some clusters contain a single observation; there's really no way that this output would generalize to other samples well:

Plotting the elbow function or cluster assignments is quick to achieve and straightforward to interpret. However, we've spoken of these techniques in terms of being heuristics. If a dataset contains a deterministic number of classes, we may not be sure that a heuristic method will deliver generalizable results.
Another drawback is that visual plot checking is a very manual technique, which makes it poorly-suited for production environments or automation. In such circumstances, it's ideal to find a code-based, automatable method. One solid option in this case is v-fold cross-validation, a widely-used validation technique.
Cross-validation is simple to undertake. To make it work, one splits the dataset into v parts. One of the parts is set aside individually as a test set. The model is trained against the training data, which is all parts except the test set. Let's try this now, again using the digits dataset:
import numpy as np from sklearn import cross_validation from sklearn.cluster import KMeans from sklearn.datasets import load_digits from sklearn.preprocessing import scale digits = load_digits() data = scale(digits.data) n_samples, n_features = data.shape n_digits = len(np.unique(digits.target)) labels = digits.target kmeans = KMeans(init='k-means++', n_clusters=n_digits, n_init=n_digits) cv = cross_validation.ShuffleSplit(n_samples, n_iter = 10, test_size = 0.4, random_state = 0) scores = cross_validation.cross_val_score(kmeans, data, labels, cv = cv, scoring = 'adjusted_rand_score') print(scores) print(sum(scores)/cv.n_iter)
This code performs some now familiar data loading and preparation and initializes the k-means clustering algorithm. It then defines cv, the cross-validation parameters. This includes specification of the number of iterations, n_iter, and the amount of data that should be used in each fold. In this case, we're using 60% of the data samples as training data and 40% as test data.
We then apply the k-means model and cv parameters that we've specified within the cross-validation scoring function and print the results as scores. Let's take a look at these scores now:
[ 0.39276606 0.49571292 0.43933243 0.53573558 0.42459285 0.55686854 0.4573401 0.49876358 0.50281585 0.4689295 ] 0.4772857426
This output gives us, in order, the adjusted Rand score for cross-validated, k-means++ clustering performed across each of the 10 folds in order. We can see that results do fluctuate between around 0.4 and 0.55; the earlier ARI score for k-means++ without PCA fell within this range (at 0.465). What we've created, then, is code that we can incorporate into our analysis in order to check the quality of our clustering automatically on an ongoing basis.
As noted earlier in this chapter, your choice of success measure is contingent on what information you already have. In most cases, you won't have access to ground truth labels from a dataset and will be obliged to use a measure such as the Silhouette Coefficient that we discussed previously.
Note
Sometimes, even using both cross-validation and visualizations won't provide a conclusive result. Especially with unfamiliar datasets, it's not unheard of to run into issues where some noise or secondary signal resolves better at a different k value than the signal you're attempting to analyze.
As with every other algorithm discussed in this book, it is imperative to understand the dataset one wishes to work with. Without this insight, it's entirely possible for even a technically correct and rigorous analysis to deliver inappropriate conclusions. Chapter 6, Text Feature Engineering will discuss principles and techniques for the inspection and preparation of unfamiliar datasets more thoroughly.
A SOM is a technique to generate topological representations of data in reduced dimensions. It is one of a number of techniques with such applications, with a better-known alternative being PCA. However, SOMs present unique opportunities, both as dimensionality reduction techniques and as a visualization format.
The SOM algorithm involves iteration over many simple operations. When applied at a smaller scale, it behaves similarly to k-means clustering (as we'll see shortly). At a larger scale, SOMs reveal the topology of complex datasets in a powerful way.
An SOM is made up of a grid (commonly rectangular or hexagonal) of nodes, where each node contains a weight vector that is of the same dimensionality as the input dataset. The nodes may be initialized randomly, but an initialization that roughly approximates the distribution of the dataset will tend to train faster.
The algorithm iterates as observations are presented as input. Iteration takes the following form:
Identifying the winning node in the current configuration—the Best Matching Unit (BMU). The BMU is identified by measuring the Euclidean distance in the data space of all the weight vectors.
The BMU is adjusted (moved) towards the input vector.
Neighboring nodes are also adjusted, usually by lesser amounts, with the magnitude of neighboring movement being dictated by a neighborhood function. (Neighborhood functions vary. In this chapter, we'll use a Gaussian neighborhood function.)
This process repeats over potentially many iterations, using sampling if appropriate, until the network converges (reaching a position where presenting a new input does not provide an opportunity to minimize loss).
A node in an SOM is not unlike that of a neural network. It typically possesses a weight vector of length equal to the dimensionality of the input dataset. This means that the topology of the input dataset can be preserved and visualized through a lower-dimensional mapping.
The code for this SOM class implementation is available in the book repository in the som.py script. For now, let's start working with the SOM algorithm in a familiar context.
As discussed previously, the SOM algorithm is iterative, being based around Euclidean distance comparisons of vectors.
This mapping tends to form a fairly readable 2D grid. In the case of the commonly-used Iris tutorial dataset, an SOM will map it out pretty cleanly:

In this diagram, the classes have been separated and also ordered spatially. The background coloring in this case is a clustering density measure. There is some minimal overlap between the blue and green classes, where the SOM performed an imperfect separation. On the Iris dataset, an SOM will tend to approach a converged solution on the order of 100 iterations, with little visible improvement after 1,000. For more complex datasets containing less clearly divisible cases, this process can take tens of thousands of iterations.
Awkwardly, there aren't implementations of the SOM algorithm within pre-existing Python packages like scikit-learn. This makes it necessary for us to use our own implementation.
The SOM code we'll be working with for this purpose is located in the associated GitHub repository. For now, let's take a look at the relevant script and get an understanding of how the code works:
import numpy as np from sklearn.datasets import load_digits from som import Som from pylab import plot,axis,show,pcolor,colorbar,bone digits = load_digits() data = digits.data labels = digits.target
At this point, we've loaded the digits dataset and identified labels as a separate set of data. Doing this will enable us to observe how the SOM algorithm separates classes when assigning them to map:
som = Som(16,16,64,sigma=1.0,learning_rate=0.5)
som.random_weights_init(data)
print("Initiating SOM.")
som.train_random(data,10000)
print("\n. SOM Processing Complete")
bone()
pcolor(som.distance_map().T)
colorbar()At this point, we have utilized a Som class that is provided in a separate file, Som.py, in the repository. This class contains the methods required to deliver the SOM algorithm we discussed earlier in the chapter. As arguments to this function, we provide the dimensions of the map (After trialing a range of options, we'll start out with 16 x 16 in this case—this grid size gave the feature map enough space to spread out while retaining some overlap between groups.) and the dimensionality of the input data. (This argument determines the length of the weight vector within the SOM's nodes.) We also provide values for sigma and learning rate.
Sigma, in this case, defines the spread of the neighborhood function. As noted previously, we're using a Gaussian neighborhood function. The appropriate value for sigma varies by grid size. For an 8 x 8 grid, we would typically want to use a value of 1.0 for Sigma, while in this case we're using 1.3 for a 16 x 16 grid. It is fairly obvious when one's value for sigma is off; if the value is too small, values tend to cluster near the center of the grid. If the values are too large, the grid typically ends up with several large, empty spaces towards the center.
The learning rate self-explanatorily defines the initial learning rate for the SOM. As the map continues to iterate, the learning rate adjusts according to the following function:

Here, t is the iteration index.
We follow up by first initializing our SOM with random weights.
Note
As with k-means clustering, this initialization method is slower than initializing based on an approximation of the data distribution. A preprocessing step similar to that employed by the k-means++ algorithm would accelerate the SOM's runtime. Our SOM runs sufficiently quickly over the digits dataset to make this optimization unnecessary for now.
Next, we set up label and color assignations for each class, so that we can distinguish classes on the plotted SOM. Following this, we iterate through each data point.
On each iteration, we plot a class-specific marker for the BMU as calculated by our SOM algorithm.
When the SOM finishes iteration, we add a U-Matrix (a colorized matrix of relative observation density) as a monochrome-scaled plot layer:
labels[labels == '0'] = 0 labels[labels == '1'] = 1 labels[labels == '2'] = 2 labels[labels == '3'] = 3 labels[labels == '4'] = 4 labels[labels == '5'] = 5 labels[labels == '6'] = 6 labels[labels == '7'] = 7 labels[labels == '8'] = 8 labels[labels == '9'] = 9 markers = ['o', 'v', '1', '3', '8', 's', 'p', 'x', 'D', '*'] colors = ["r", "g", "b", "y", "c", (0,0.1,0.8), (1,0.5,0), (1,1,0.3), "m", (0.4,0.6,0)] for cnt,xx in enumerate(data): w = som.winner(xx) plot(w[0]+.5,w[1]+.5,markers[labels[cnt]], markerfacecolor='None', markeredgecolor=colors[labels[cnt]], markersize=12, markeredgewidth=2) axis([0,som.weights.shape[0],0,som.weights.shape[1]]) show()
This code generates a plot similar to the following:

This code delivers a 16 x 16 node SOM plot. As we can see, the map has done a reasonably good job of separating each cluster into topologically distinct areas of the map. Certain classes (particularly the digits five in cyan circles and nine in green stars) have been located over multiple parts of the SOM space. For the most part, though, each class occupies a distinct region and it's fair to say that the SOM has been reasonably effective. The U-Matrix shows that regions with a high density of points are co-habited by data from multiple classes. This isn't really a surprise as we saw similar results with k-means and PCA plotting.
Victor Powell and Lewis Lehe provide a fantastic interactive, visual explanation of PCA at http://setosa.io/ev/principal-component-analysis/, this is ideal for readers who are new to the core concepts of PCA or who are not quite getting it.
For a lengthier and more mathematically-involved treatment of PCA, touching on underlying matrix transformations, Jonathon Shlens from Google research provides a clear and thorough explanation at http://arxiv.org/abs/1404.1100.
For a thorough worked example that translates Jonathon's description into clear Python code, consider Sebastian Raschka's demonstration using the Iris dataset at http://sebastianraschka.com/Articles/2015_pca_in_3_steps.html.
Finally, consider the sklearn documentation for more details on arguments to the PCA class at http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html.
For a lively and expert treatment of k-means, including detailed investigations of the conditions that cause it to fail, and potential alternatives in such cases, consider David Robinson's fantastic blog, variance explained at http://varianceexplained.org/r/kmeans-free-lunch/.
A specific discussion of the Elbow method is provided by Rick Gove at https://bl.ocks.org/rpgove/0060ff3b656618e9136b.
Finally, consider sklearn's documentation for another view on unsupervised learning algorithms, including k-means at http://scikit-learn.org/stable/tutorial/statistical_inference/unsupervised_learning.html.
Much of the existing material on Kohonen's SOM is either rather old, very high-level, or formally expressed. A decent alternative to the description in this book is provided by John Bullinaria at http://www.cs.bham.ac.uk/~jxb/NN/l16.pdf.
For readers interested in a deeper understanding of the underlying mathematics, I'd recommend reading the work of Tuevo Kohonen directly. The 2012 edition of self-organising maps is a great place to start.
The concept of multicollinearity, referenced in the chapter, is given a clear explanation for the unfamiliar at https://onlinecourses.science.psu.edu/stat501/node/344.
In this chapter, we've reviewed three techniques with a broad range of applications for preprocessing and dimensionality reduction. In doing so, you learned a lot about an unfamiliar dataset.
We started out by applying PCA, a widely-utilized dimensionality reduction technique, to help us understand and visualize a high-dimensional dataset. We then followed up by clustering the data using k-means clustering, identifying means of improving and measuring our k-means analysis through performance metrics, the elbow method, and cross-validation. We found that k-means on the digits dataset, taken as is, didn't deliver exceptional results. This was due to class overlap that we spotted through PCA. We overcame this weakness by applying PCA as a preprocess to improve our subsequent clustering results.
Finally, we developed an SOM algorithm that delivered a cleaner separation of the digit classes than PCA.
Having learned some key basics around unsupervised learning techniques and analytical methodology, let's dive into the use of some more powerful unsupervised learning algorithms.




















 Download code from GitHub
Download code from GitHub

